When we think of the word marker, the first thing that comes to our minds is something that is used to indicate a place. For example, it can be your current location on Google Maps or it can be the place where you planted some seeds in your garden. Similarly, in genomics studies, we can find marker genes in bacterial genomes. In this article, I will introduce you to marker genes used in metagenomics analysis, how they are used and walk you through an example of a commonly used gene prediction tool.
What are Marker Genes/Genetic Markers?
According to Wikipedia,
A genetic marker is a gene or DNA sequence with a known location on a chromosome that can be used to identify individuals or species.
As a result of mutations and alterations within the genome, these genes can vary depending on their composition and location.
What are Single-Copy Marker Genes?
In bacterial cells, single-copy marker genes are expected to occur once. In other words, each bacterial cell contains only one copy of each of these single-copy marker genes. These genes are essential for the life-functions and can be found in the majority of the bacterial species.
Previous efforts have been made to identify marker genes that can resolve closely related organisms. Protein-coding marker genes which are rarely horizontally transferred and exist in single copies within genomes have been identified [1]. These include a set of 40 marker genes [2,3] and 107 marker genes [3].
Usage of Marker Genes
Marker genes are commonly used in taxonomic profiling of environmental samples to identify gene families. These genes are also used in phylogenetic inference to reconstruct the evolutionary history of organisms.
Recently, reference-free binning tools such as MaxBin and SolidBin have used single-copy marker genes to identify the number of species in a given sample. Moreover, tools such as MyCC use single-copy marker genes to refine resulting clusters.
Gene Predictors
Gene predictors can be used to extract marker genes. Some popular gene prediction tools include,
Example Usage of FragGeneScan
Let us see how we can use FragGeneScan to predict genes. Firstly, you can download FragGeneScan from
- SourceForge: https://sourceforge.net/projects/fraggenescan/files/latest/download
- GitHub: https://github.com/gaberoo/FragGeneScan
You can follow the instructions provided in the README file to compile and run FragGeneScan.
You can see the following parameters and options of FragGeneScan.
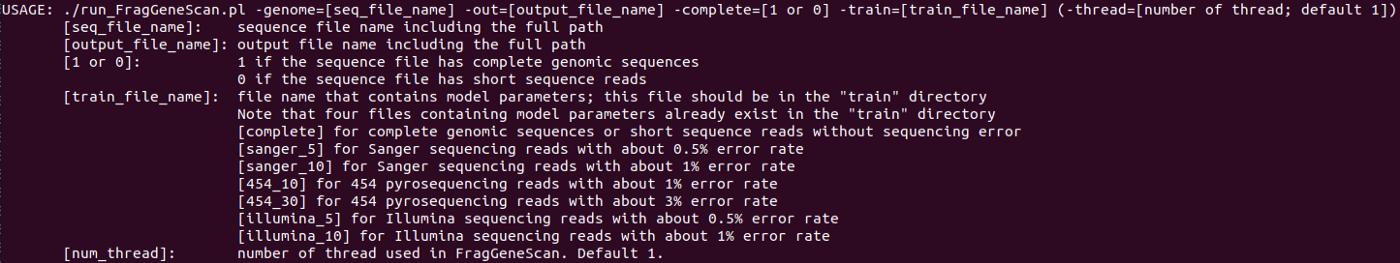
If you have a complete genomic sequence, you can run FragGeneScan to predict its genes as follows.
./run_FragGeneScan.pl -genome=<sequence_file> -out=<output_file> -complete=1 -train=complete -thread=<num>If you have a set of assembled contigs, you can run FragGeneScan to predict its genes as follows.
./run_FragGeneScan.pl -genome=<contigs_file> -out=<output_file> -complete=0 -train=complete -thread=<num>FragGeneScan generates four files with their contents as follows.
<output_file>.out: coordinates of putative genes<output_file>.fnn: nucleotide sequences corresponding to the putative genes in<output_file>.out<output_file>.faa: amino acid sequences corresponding to the putative genes in<output_file>.out<output_file>.gff: gene prediction results
Once you have obtained these files, you can use the <output_file>.faa file along with HMMER to determine the single-copy marker genes in the sequences.
Final Thoughts
Marker genes have become a very powerful aspect in bioinformatics research which has allowed researchers to gain insights into the taxonomic information and evolutionary history of bacterial and archaeal species. The field of metagenomics benefits immensely from studies based on marker genes.
I hope you found this article useful. Feel free to try out the tools mentioned in this article and play around with examples.
Cheers, and stay safe!
References
[1] Microbial abundance, activity and population genomic profiling with mOTUs2 (https://motu-tool.org/fetchMG.html)
[2] Ciccarelli et al. (2006) Toward Automatic Reconstruction of a Highly Resolved Tree of Life, SCIENCE 03: 1283–1287
[3] Wu D, Jospin G, Eisen JA (2013) Systematic Identification of Gene Families for Use as “Markers” for Phylogenetic and Phylogeny-Driven Ecological Studies of Bacteria and Archaea and Their Major Subgroups. PLoS ONE 8(10): e77033. https://doi.org/10.1371/journal.pone.0077033
This article was originally published in The Computational Biology Magazine on Medium.
Cover image by Mahmoud Ahmed from Pixabay
You can find the original article at https://medium.com/computational-biology/marker-genes-and-gene-prediction-of-bacteria-5fa4cb7802f3

