With the development of various methods to obtain data from living beings, there has been an explosion in biological data which is readily available to be used. However, such vast amounts of data will be of no use if there is no proper way to execute a series of steps to manipulate the data as we want, to output desired results. This is where Workflow Management Systems come in handy.
Before diving further, let me explain today’s outline. I will be walking you through the following sections in this article.
- What is a Workflow?
- What is a Workflow Management System?
- What is a Bioiformatics Workflow Management System?
- Unipro UGENE
I won’t be going deep on how to create bioinformatics workflows, but I will explain about workflow management systems and introduce you to Taverna, a workflow management system widely used by bioinformaticians.
What is a Workflow?
A workflow consists of a set of activities which are enabled by the systematic organization of resources that transform materials, provide services, or process information. It can be depicted as a sequence of operations to complete a process.
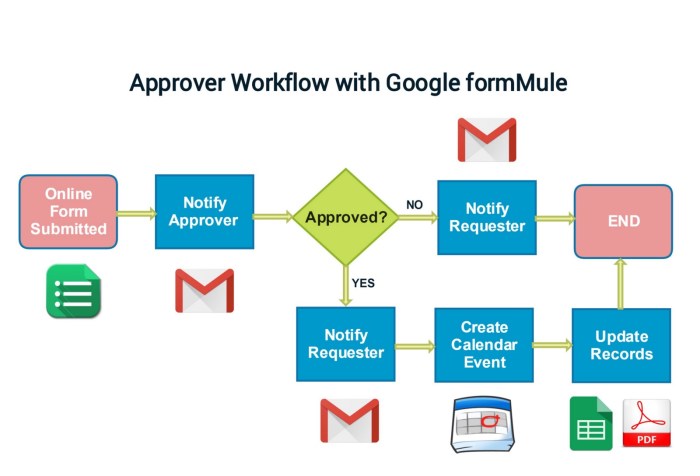 Example Workflow (Image Source: YouTube)
Example Workflow (Image Source: YouTube)
What is a Workflow Management System?
A Workflow Management System (WMS) is a software that provides an infrastructure to setup, execute, and monitor scientific workflows.
Workflow management systems emerged as an answer to the problem of how to carry out and automate complex processes on larger volumes of heterogenous data. They visualize workflows in the form of workflow diagrams, depicting inputs, outputs, services and data flows. They also allow to save workflows for publishing and sharing.
What is a Bioinformatics Workflow Management System?
A bioinformatics workflow management system is a specialized form of workflow management system designed specifically to compose and execute a series of computational or data manipulation steps that is related to bioinformatics.
Such systems show an abstract representation of the computation and how it proceeds in the form of a directed graph. Each node of the graph represents a task to be executed and each edge represents either data flow or execution dependency between different tasks. The system provides a visual front-end, which allows the user to build and modify different workflows with little or no programming expertise.
Some famous bioinformatics workflow management systems include,
 Taverna Workbench (Image Source: Apache Taverna)
Taverna Workbench (Image Source: Apache Taverna)

Galaxy (Image Source: MGEscan)

Unipro UGENE
Unipro UGENE is a free open-source cross-platform bioinformatics software. It allows you to View, edit, annotate and align DNA, RNA and protein sequences, work with 3D structures and surface algorithms and model workflows using the Workflow Designer. You can read more details about UGENE from here.

UGENE Loading Screen
You can download Unipro UGENE from here. Once you have downloaded the installer, follow the steps given in the readme.txt before installing. Make sure to copy the sample folder provided in the download containing sample files in different formats.
Once installed and the application is opened, you will see the Start Page as depicted in Figure 1. (I have opened one file before and you can see it under recent files. If you open UGENE for the first time, you will not see any files here.)
 Figure 1
Figure 1Building a simple workflow using UGENE Workflow Designer
UGENE Workflow Designer allows to create and run complex computational workflows even if he or she is not familiar with any programming language.
We will start by building a simple workflow; find a given pattern in a sequence or in sequences and save it as annotations.
Click on Create Workflow option in the Start Page. You will see the workflow design area as shown in Figure 2.
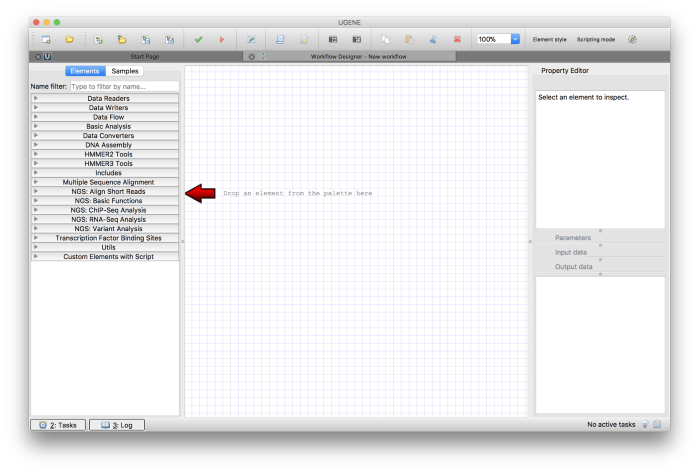 Figure 2
Figure 2
Drag and drop Find Pattern element from Basic Analysis section in the list of elements as shown in Figure 3.
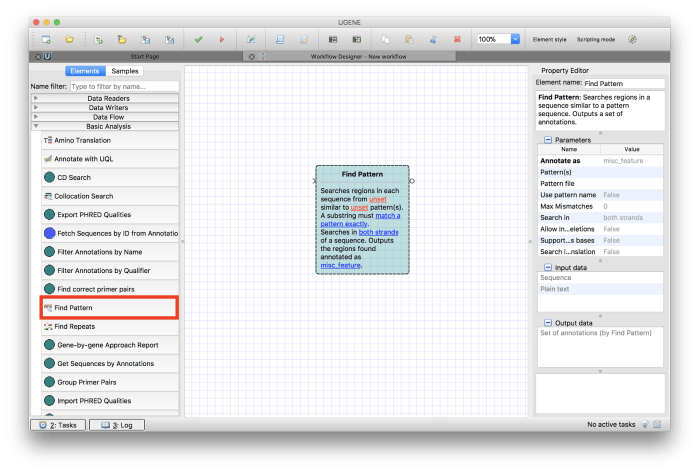 Figure 3
Figure 3
Now add Read Sequence component from Data Readers section in the list of elements as shown in Figure 4.
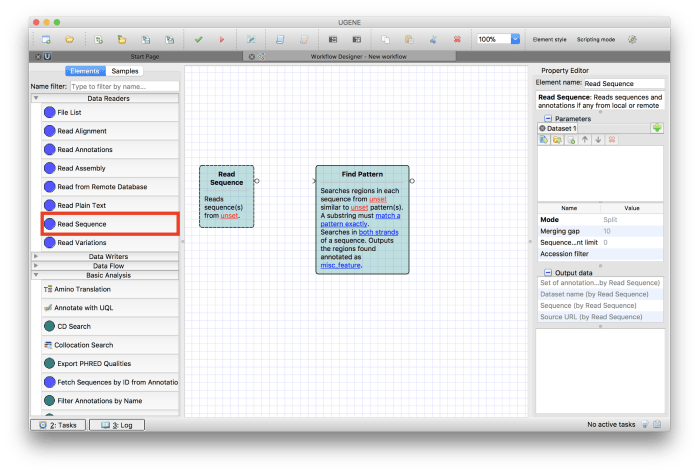 Figure 4
Figure 4
Click on the loop mark on Read Sequence component, drag it on the half loop mark on Find Pattern element to connect them as shown in Figure 5.
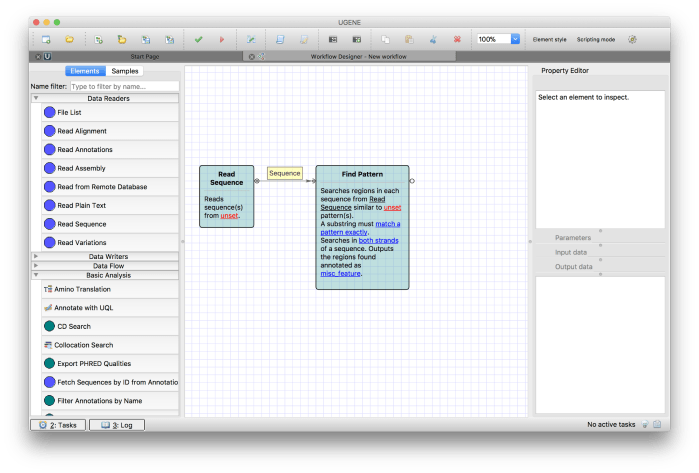 Figure 5
Figure 5
Now similarly, add Write Sequence component from Data Writers section in the list of elements and connect it to Find Pattern element. Final workflow should look as given in Figure 6.
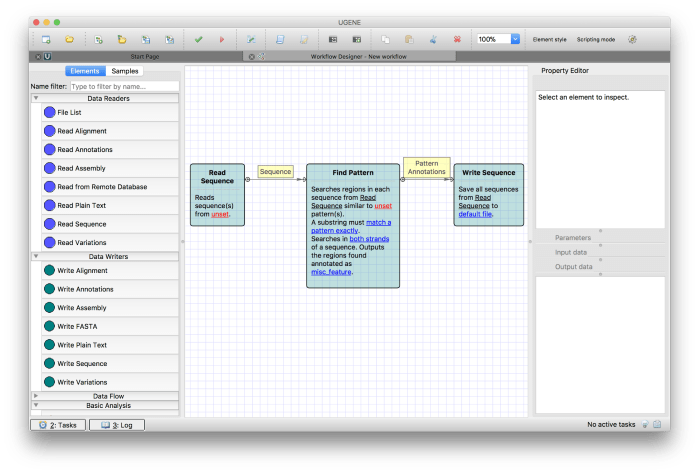 Figure 6
Figure 6
Now we have to define the file containing the input sequences. In order to do so, click on the word unset in the Read Sequence component and you will see its details in the Property Editor on the right-hand side of the screen. Under Dataset1, click on Add files icon. Refer Figure 7.
 Figure 7
Figure 7
You will see that the sample folder is opened. For this example we will be selecting the following file in the given path. Refer Figure 8.
samples → Genebank → murine.gb
 Figure 8
Figure 8Now we have to define the pattern we want to find. Click on the word unset in the Find Pattern component and you will see its details in the Property Editor on the right-hand side of the screen. Under Parameters, enter the pattern you want to match as the value for Pattern(s) under Annotate as section. Refer Figure 9. I have used the following pattern.
TTCCGAGGGACACTAGGCTGACTCCATC
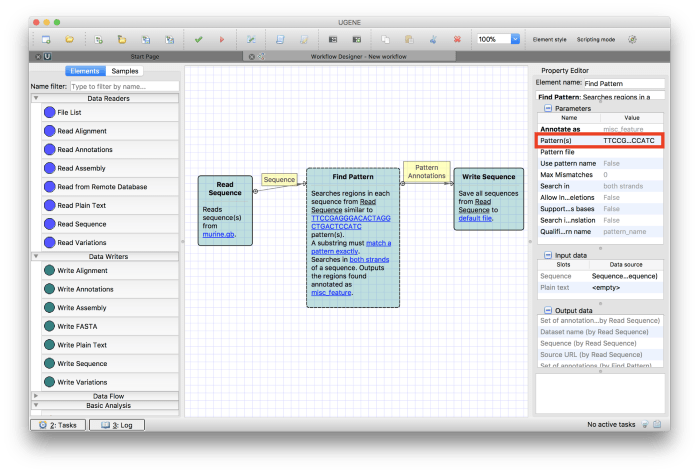
Our final step is to define the output file. Click on default file in the Write Sequence component and you will see its details in the Property Editor on the right-hand side of the screen. Under Parameters, enter the output file name as pattern.gb to Output file value. Change the Document format of the output file to GeneBank. Final values should be as in Figure 10.
 Figure 10
Figure 10
Now you can run the workflow by clicking on the Run workflow button in the tool bar as shown in Figure 11.
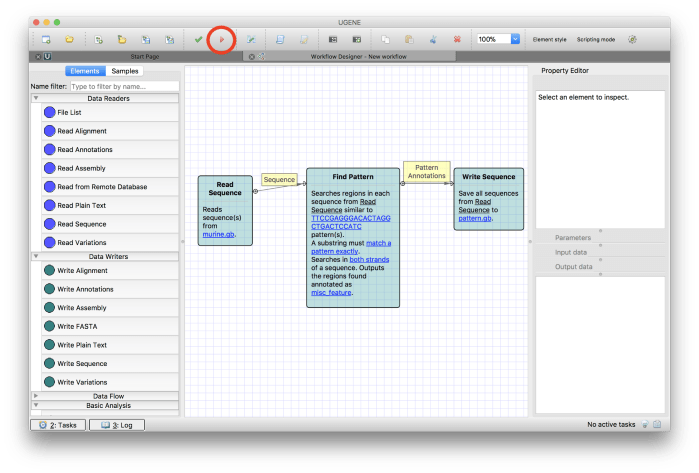 Figure 11
Figure 11
A report including all the information about the workflow execution will be shown as in Figures 12 and 13.
 Figure 12
Figure 12
 Figure 13
Figure 13
You can open the results file (pattern.gb) and view it as in Figure 14.
 Figure 14
Figure 14
You can play around with the tool and see other available components, create workflows and run them.
Hope you got a basic idea about bioinformatics workflow management software to start experimenting with.
Cheers! 😊
Originally published in Medium.com publication Towards Data Science.

